Imaging platform
Institute of Molecular Biology and Pathology

Visualizing cell dynamics
The IBPM microscopy platform hosts last-generation widefield light microscopy equipment and advanced imaging software for automated image analyses.
It supports versatile applications for both fixed and live cell samples and provides opportunities for coupling dynamic studies (time-lapse recording) with high resolution qualitative and quantitative image analysis.
The Center is open to external users, click here for access rules.
Applications
- Time-lapse recording of cellular processes in live cells over days
- Detecting intracellular molecular interactions
- Reconstructing cellular and subcellular structures at 3D level
- Quantitative analysis of imaging data
About us
The platform was first established in 2009 with support from Fondazione Monte Paschi, Fondazione Roma and Assicurazioni Generali S.p.A. It has since been supported by two dedicated grants from Regione Lazio and by the CNR Project “Impara - Imaging from Molecules to Preclinical Studies” funded by the Ministry of Education and Research for the development of National research infrastructures. The platform is hosted at the Department of Biology and Biotechnology “Charles Darwin” of Sapienza University.
Latest News
New Application Note: Capture More, Damage Less: Spinning Disk Confocal Imaging for Long-Term Live Studies
Long-term live-cell imaging is a technique for the study of dynamic biological processes with high temporal and spatial resolution. This application note presents the benefits of using the X-Light V3 spinning disk confocal system from CrestOptics for extended time-lapse imaging of live samples.
Case studies: Autophagy monitoring and Analysis of cell division
"Plenty of fish in the sea" selected image in Biologists@100 conference, JCS - Image Competition
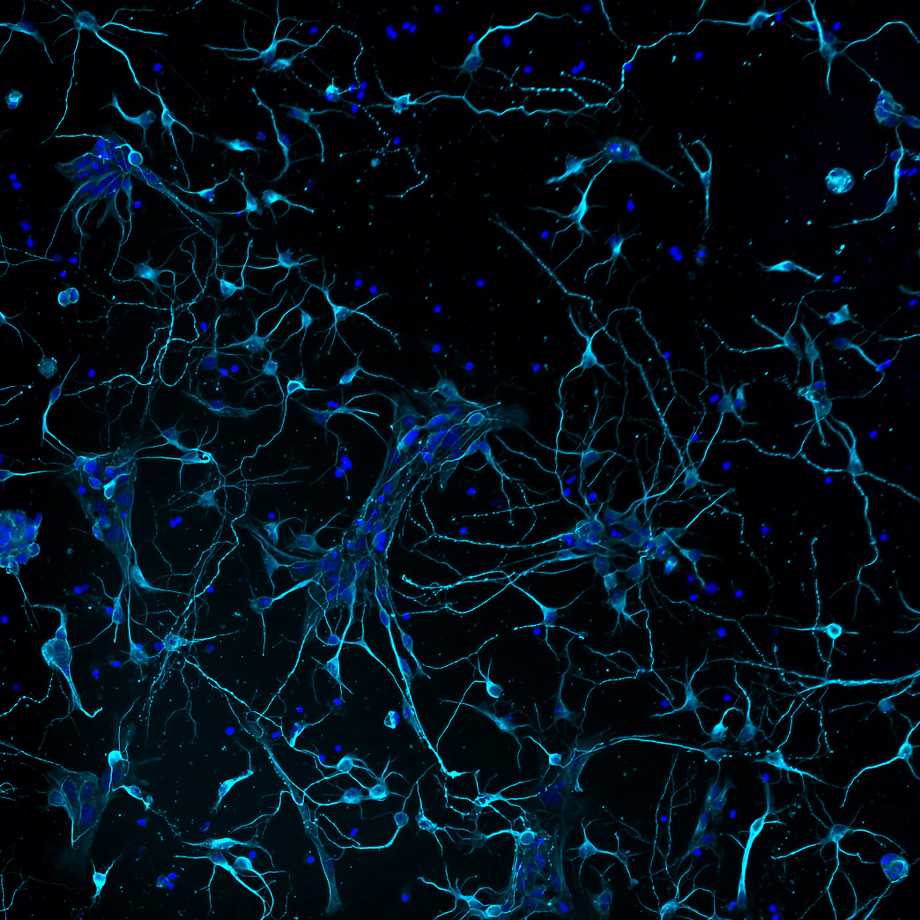
Plenty of fish in the sea - Ludovica Altieri
Murine primary cortical neurons developing interconnections, stained with neuronal tubulin (cyan) and DAPI (blue). This image has been selected among the 15 finalists of the competition organised by The Node and FocalPlane (The Company of Biologists) and was presented at Biologists@100 gallery in Liverpool.
Imaged on a Nikon microscope implemented with a CrestOptics confocal spinning disk module with post-processing using NIS Elements AR by Nikon. Acquired at the IBPM Institute of Molecular Biology and Pathology – CNR National Research Council of Italy, c/o Department of Biology and Biotechnology “Charles Darwin” – Sapienza University of Rome.
Members
![]() Lia Asteriti
Lia Asteritilia.asteriti@uniroma1.it
![]() Francesca Degrassi
Francesca Degrassifrancesca.degrassi@uniroma1.it
![]() Giulia Fianco
Giulia Fiancogiulia.fianco@gmail.com
![]() Giulia Guarguaglini
Giulia Guarguaglinigiulia.guarguaglini@uniroma1.it
![]() Patrizia Lavia
Patrizia Laviapatrizia.lavia@uniroma1.it
![]() Valerio Licursi
Valerio Licursivalerio.licursi@uniroma1.it
![]() Federica Polverino
Federica Polverinofedericapolverino@cnr.it
![]() Davide Valente
Davide Valentedavide.valente@cnr.it
![]() Ludovica Altieri
Ludovica Altieriludovicaaltieri@cnr.it
Sponsors